Difference between revisions of "Future version"
| Line 14: | Line 14: | ||
NMReDATA is being integrated with [[Jmol | Jmol/JspecView ]] | NMReDATA is being integrated with [[Jmol | Jmol/JspecView ]] | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
= Comparison of assigned data= | = Comparison of assigned data= | ||
Revision as of 09:23, 23 October 2019
This page contains item for future releases. The next planned release is 2.0. Currently we aim for the 2019 Symposium to decide about features. Some may be pushed to later releases. All item on the list are currently in equal consideration, no matter if they are on separate pages or not. The discussion pages can be used for discussions.
All of this has to be refined - comments are welcome!
Contents
- 1 3D Structures
- 2 Databse policy
- 3 Jmol
- 4 Comparison of assigned data
- 5 Certification
- 6 Role(s) and scope of the “assignment records”
- 7 SDF files generated from experimental data
- 8 SDF files generated from calculated data
- 9 SDF files generated from literature data
- 10 SDF files generated after revision of existing SDF files
- 11 Concerning symmetry
- 12 Why not add more data in NMReDATA tags?
- 13 License
- 14 Author
- 15 Solvent
- 16 nmrML
3D Structures
Discussion on New rules for the inclusion 3D structure
Databse policy
Discussion on NMReDATA database
Jmol
NMReDATA is being integrated with Jmol/JspecView
Comparison of assigned data
This is a working proposition
A tag comparing the data in the current file with those from other files (with the same labels for the assignment) > <NMREDATA_COMPARISON>
Externaldata1=... ref to an external .sdf file from the current or external record. Externaldata2=... H1 1.54 1.50 1.66 (the first value is the reference value in NMREDATA_ASSIGNMENT tag of the current file the second from Externaldata1, etc.) H2 1.54 1.50 1.66 Chi2 0.3 0.5 (these could be the chi square of the reference with the external data )
Coupling could also be compared...
> <NMREDATA_COMPARISON>
Externaldata1=... ref to an external .sdf file from the current or external record. Externaldata2=... J(H1,H2) 1.54 1.50 1.66 (the first value is the reference value in NMREDATA_ASSIGNMENT tag of the current file the second from Externaldata1, etc.) J(H1,H3) 1.54 1.50 1.66 Chi2 0.3 0.5 (these could be the chi square of the reference with the external data)
Other comparisons could be made, for example from signal intensities in HSQC spectra
> <NMREDATA_COMPARISON>
Externaldata1=... ref to an external .sdf file from the current or external record. Externaldata2=... I(NMREDATA_13C_1J_1H,HC1,H1) 122 154 143 (the first value is the reference value in NMREDATA_ASSIGNMENT tag of the current file the second from Externaldata1, etc.) I(NMREDATA_13C_1J_1H,HC2,H2) 132 151 163 Chi2 5.3 3.5 (these could be the chi squared)
Certification
As we have seen, software can judge the consistency of spectral data and their agreement with a proposed structure. The algorithms implemented for this can vary and are not the subject of this paper. We show how the evaluation of an assignment, as recorded in an NMReDATA, can be included in the file in a manner which prevents manipulation of the data. We call this certification of NMReDATA. The NMREDATA\_CERTIFICATION tag is used for recording certifications. In \cite{paper1} the tag was mentioned, but not fully specified. This is done here as part of NMReDATA 1.2.
We use public-key cryptography in order to achieve this. The main advantage of this method is that it is possible to check the authenticity with only the public key of the certifier, which can be easily distributed using existing infrastructures like key servers. It would also be possible to store the certification on a server, but this would make the future verification of the certification depend on the availability of server storage.
We demonstrate the use of the NMREDATA\_CERTIFICATION tag with an example, shown in Fig.~\ref{fig:certification}. It contains one certification, further certification could be added, in a separate NMREDATA\_CERTIFICATION_X tag. \texttt{Certificate=} points to a certificate in a separate tag, which contains those parts of the document which are certified, as seen by the software, signed and compressed by an encryption software compatible with the OpenPGP standard \cite{OpenPGP} (e.g.\ GnuPG), in ascii armor format. The separate tag is needed because of problem with line breaks (\ is desired in NMReDATA tags, but in the certificate we cannot have them) Used_tags tells the tags included in the certification. The certified document can contain only those parts of the original document, in particular only those spectra used. It must contain the other entries (in this case \texttt{Confidence\_level=} and \texttt{Quality\_level=}) in the certification. The technical process of generating and reading the certificate will be explained in the next paragraph. The entries in a certification, apart from \texttt{Software=} and \texttt{Certificate=}, are decided by the authors of the certification software. The \texttt{Author=} entry is recommended, and refers to the author(s) of the software. In case of nmrshiftdb2 the software gives a confidence level and a quality level as a measurement for the overall assignment.
Other software might have its own criteria. For example, a computer-aided structure elucidation (CASE) software might be able to decide if a structure is a unique solution and therefore add an entry like \texttt{Unique\_solution=YES} to the certification.
> <NMREDATA_CERTIFICATION_NMRSHIFTDB2>
Software=nmrshiftdb2
Version=1.0 \hl{DJ:I propose to add a verion number}
Author=Stefan Kuhn and the nmrshiftdb2 team
Confidence_level=good
Quality_level=4/10, revision
Used_tags= NMREDATA_CERTIFICATION_NMRSHIFTDB2 , NMREDATA_INCHI, NMREDATA_ASSIGNMENT
Certificate=NMRSHIFTDB2_CERTIFICATE
> <NMRSHIFTDB2_CERTIFICATE>
-----BEGIN PGP MESSAGE-----
Version: GnuPG v2.0.22 (GNU/Linux)
owHtWW1oW1UYbmq30eKYgkNZtcbYybqedfec+5VLHaPrsnajDbFNcSpYk+42yZom
...
7W+OdV3c33V4/YYPK79aPv3EkasP70zccfm5E/8A
=/r/c
-----END PGP MESSAGE-----
In order to do certifications, the certifying authority must generate a key pair, using any OpenPGP-compatible software. The public key should be available for download. We show the procedure here using GnuPGP, but any compatible software can be used for the certificiation as well for the verification. The process is of course performed automatically by the software. Firstly, the quality parameters are calculated. In case of nmrshiftdb2 these are the quality and confidence (these values are also shown if a spectrum is submitted to nmrshiftdb2 or the Quick Check on www.nmrshiftdb.org is used, see Figure~\ref{fig:qc}). These are then added to the NMRedata file to certify, so that the NMREDATA\_CERTIFICATION tag is complete except the \texttt{Certificate=} entry and the rest of the file is unchanged. OpenPGP is then used to do the signing: <dschober@ipb-halle.de> 2018-09-04T16:17:42.041Z: certifying authority I think there needs to be a disambuguation between 'trusted data' and validation. Many readers will not be familiar with 'certification'. Also state more clearly
gpg --output example.nmredata.sdf.sig --armor --sign example.nmredata.sdf
Notice we use the --armor option to create an ascii output. The resulting signed file is then put into the NMReDATA file as in Fig.~\ref{fig:certification}. In order to verify this the signature can be decrypted using this command:
gpg --output example.nmredata.sdf --decrypt example.nmredata.sdf.sig
The resulting file should be the same as the one which was signed, including the NMREDATA\_CERTIFICATION entry and the quality and confidence levels. The public key, which is available on keyservers and on www.nmrshiftdb.org, must be installed for the verification. GnuPGP will also print a message saying \texttt{gpg: Good signature from "nmrshiftdb2 (Key for NMReDATA certification) <nmrshiftdb2-admin@uni-koeln.de>"}, which tells the user the file has been properly signed by nmrshiftdb2. The signing cannot be reproduced by a third party and no change to the signed content are possible.
Role(s) and scope of the “assignment records”
The NMR record can be generated from experimental data (this is how the format was designed), but data may also originate from simulations, predictions, etc.
Tools to compare, evaluate, validate, and check consistency of “assignment records” will certainly be developed.
Assignment records can be generated by commercial software, but also by diverse tools analysing NMR data, homemade processing tools, simulation software, etc. This is why it is important to have a format of data including a maximum of options to be as flexible as possible, even if not all possible uses are clearly defined and used immediately. Ideally, the .sdf files should be converted into other file formats or spectral description without loss.
Multiple records
We should see as an advantage if databases include multiple "assignment records" associated to the same molecule or the same set of NMR spectra. Some could be old, originating from incomplete literature data. Others could include errors because they originate from bulk data processed automatically. But finally, a computer could verify or create a robustly validated record combining all the other data. Aggregated record could be generated by NMR software/database scoring available data for consistency, calculated chemical shifts and spectral simulations. They could refine chemical shifts and couplings, etc.
SDF files generated from experimental data
When the NMR data originate from experimental spectra, they may be quite crude (simple automated integration, peak-picking). At the other extreme, the data may follow complex automated or careful and manual expert analysis. The NMReDATA has the flexibility to code diverse quality of data: They may be partial, incomplete, contain inconsistencies, impossible features, etc.
Here is a (non-comprehensive) list of possible situations with respect to NMReDATA:
- only 1D 1H NMR data (with or without integration, coupling, etc.).
- only 1D 13C data (just from a simple peak peaking)
- only 1D data but for multiple isotopes
- full analysis based on computer-assisted software (such as ACDLabs Structure Elucidator, Bruker CMC-se or Mestrelab Mnova) or web platform (such as cheminfo.org).
- 1D and 2D data processed automatically with ambiguities on the signal assignment and partial (for example not all signals are assigned) and/or ambiguous (due to lack of resolution, or other problems)
- The file may not contain the actual assignment, only the structure and the list of chemical shift (the assignment could be added by NMR tools).
- The data may come from a scientific report i.e. the text providing the description of the spectra.
Scripts could be written to convert such a "pure text" description into .sdf file and include the .mol file.
For assignment work made with only "paper and pencil", a tool allowing to draw a molecule, enter lists of signal names and 2D correlations could be easily made. We could consider to accept .pdf or pictures of the spectra when the original files do not exist anymore.
SDF files generated from calculated data
The NMR data may originate from DFT calculations or any other type of predictor of chemical shifts, and/or coupling. In such a case, a general tag is added to provide information about the software. For example:
> <NMREDATA_ORIGIN> Source=Calculation method=DFT Geometry=method/basis set Shielding=method_basis set Coupling=method_basis set Software=... Version=...
References to addition files located in a folder called "DFT_GIAO_calculations" could be added. They could include: -the equation used to convert sheelding in chemical shifts (for 1H, 13C)
csH=((Sxx+Syy+Szz)/3-60)*0.98+5" csC=((Sxx+Syy+Szz)/3-130)*1.1+3.1" JHH=((???)/3-130)*1.1+3.1"
-the list of conformers calculated, their energies ?, the Boltzman ratio ?
-all 3D structures of the conformations in a conformations.sdf file (not the main NMReDATA .sdf file). The main .sdf file can contain the 3D structure of the lowest energy conformation and the flat structure only.
-all the output files of the shielding (and coupling) calculations ? Also the geometry optimization ?
Examples (tentative) of .nmredata.sdf files originating from Gaussian can be found here: androsten data
SDF files generated from literature data
This is a working proposition under review
When the NMR data originate from publications, a reference to the published paper/book/thesis is given in the NMREDATA_LITERATURE tag.
> <NMREDATA_LITERATURE> Source=Journal DOI=DOI_HERE (if Reference field is DOI specify it here) CompoundNumber=label used in the reference to designate the compound (typically a number in boldface)
> <NMREDATA_LITERATURE> Source=Book ISBN=ISBN_HERE (if Reference field is DOI specify it here) CompoundNumber=label used in the reference to designate the compound (typically a number in boldface)
> <NMREDATA_LITERATURE> Source=Thesis Thesis=HTML link here (if available if not "LastName, Firstname(s), institution providing the degree, city, country, year of publication. CompoundNumber=label used in the reference to designate the compound (typically a number in boldface)
SDF files generated after revision of existing SDF files
Assignment records may be generated after revision from experimental, literature, prediction data, etc. Ideally, the original .sdf files should also be generated to facilitate comparison or exist somewhere and be referred to. In both cases reference should be given.
> <NMREDATA_UPDATE> Source=Record Record_number=ref_to_the_original_record (multiple reference is allowed for aggregation of records – separated by “,”). Date =date.... standard format for date Correction="fixed assignments of C(13) and C(15)"
This is also to be refined according to future developments.
Concerning symmetry
Magnetic non-equivalence
For symmetrical molecules a difficulty may arise to code coupling and 2D correlations.
Reminder: Couplings are not directly associated to atoms, but to labels (in the NMREDATA_ASSIGNMENT tag). Labels are associated to one or more atoms (in case of symmetry/fast rotation, etc.).
Examples of difficulty/solution concerning scalar coupling: For the 1H spectrum of 1, 2 dichlorobenzene, we have two multiplets in the 1D 1H spectrum (two different protons in an AA’XX’ system) so if the SDF file includes two labels (one for A and one for X, each pointing to two atoms), in principle one can only give one coupling: the JA,X (no JA,A or JA,X'). But if one desires to specify all the couplings, give two different "labels" to A and A' (each pointing to only one atom), so that different coupling can be given for JA,X, JA',X,JA,A', JX,X'. This may be desired so that the 1D spectrum can be simulated with the correct non-equivalence effect.
HMBC correlations in symmetrical molecules
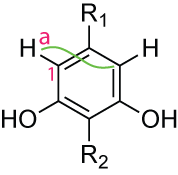
Consider the following molecules:
A 3JC,H HMBC correlation will be visible between the proton a and C(1) that seems to be the directly bound carbon. Because the carbons 1J and 3J bond, relative to a proton are symmetrical. Software may see the correlation as 1J, but, it should be able to analyse the NMREDATA_ASSIGNMENT tag and see that a and C(1) are pointing to two atoms, and that the correlation may correspond to any combination of the four possible pairs. Two pairs will seem as the actual 3J and two as the 1J.
Why not add more data in NMReDATA tags?
We consider that our task is to focus on NMR data. But SDF files could (and probably should!) also include other experimental data such as:
1) The origin of the molecule. This may include the extraction method and the plant it originates from, in phytochemistry, or the reaction producing it.
2) MS data
3) other spectral data
In principle authors can add any tag provided they have tools to do it and requests from the journals... such data could have the following form...
The software producing SDF files including NMReDATA, should read SDF files and write SDF files only adding (or modifying/reviewing) the NMReDATA data. Any other SDF tags should be passed from the file which is read to the file which is generated.
Important note on tag names: To comply with the SDF format specification, tag names cannot contain hyphen (-), period (.), less than (<), greater than (>), equal sign (=), percent sign (%) or blank space ( ). Tag names must begin with an alpha character and can contain alpha and numeric characters after that, including underscore.
License
The NMReDATA\_LICENCE is intended to give a license under which the author(s) of the document have published it. Since the number of potential licenses is not limited, we do not restrict the choice here. The license should be easily identifiable and ideally a link to the license text should be provided as well. There can be only one license for a document and its different versions. If a change of license is required, a new document is required. It is up to the authors to decide which parts of the old document they may reuse, depending on the legal situation and previous license. The format does not imply any legal aspects and users need to judge the legal situation for themselves.
Author
The AUTHOR tag records authors of the NMReDATA file. We have decided to restrict the entries to names, ORCIDs, and affiliations of authors. Since documents can potentially be edited, authors can be given for different versions of the document. The versions are numbered and ordered by integers. For each version, any number of authors can be given. So within the AUTHOR tag, there can be any number of \texttt{Version=} entries. Below each of these, there can be any number of Author= entries. Below each author there can be an \texttt{Affiliation=} and \texttt{ORCID=} entry. Fig.~\ref{fig:author} gives an example of document, which had one original author and was then changed by two authors to derive a second version.
> <AUTHOR> Version=1 Author=Stefan Kuhn Affilition=De Montfort University, Leicester ORCID=0000-0002-5990-4157 Version=2 Author=John Doe Affilition=University of Nowhere, Fictionland Author=Joe Bloggs
Solvent
We also introduce a more detailed specification of the solvent. So far, this was a text field. If wished, solvents, buffers, and references can be specified exactly. For each of them <compound name>,<concentration>,<concentration unit> are given in this order. Several lines are possible in case of mixtures. Fig.~\ref{fig:solvent} gives an example of a structured NMREDATA\_SOLVENT. Note that there is an additional entry for Cytocide, such additions, following the same format, are possible. If a solvent entry does not start with \texttt{Solvent=} it is assumed to be a simple solvent entry with not structure.
> <NMREDATA_SOLVENT> Solvent=D2O, 100, % Buffer=sodium phosphate, 50, mM Cytocide=sodium azide, 500, uM Reference=DSS, 0.1,%,
nmrML
NMReDATA works in combination with the nmrML standard \cite{pmid29035042}. Since nmrML is a standard for raw data, whereas NMReDATA is dealing with processed data and higher level concepts like peaks or couplings, they complement each other. NMReDATA allows the use of nmrML data in the NMR record instead of the vendor raw data.